etf <- etf_vix[1:55, 1:3]
# Split-------------------------------
h <- 5
etf_eval <- divide_ts(etf, h)
etf_train <- etf_eval$train
etf_test <- etf_eval$testBayesian VAR and VHAR
var_bayes() and vhar_bayes() fit BVAR and
BVHAR each with various priors.
-
y: Multivariate time series data. It should be data frame or matrix, which means that every column is numeric. Each column indicates variable, i.e. it sould be wide format. -
porhar: VAR lag, or order of VHAR -
num_chains: Number of chains- If OpenMP is enabled, parallel loop will be run.
-
num_iter: Total number of iterations -
num_burn: Number of burn-in -
thinning: Thinning -
coef_spec: Coefficient prior specification.- Minneosta prior
- BVAR:
set_bvar() - BVHAR:
set_bvhar()andset_weight_bvhar() - Can induce prior on
using
lambda = set_lambda()
- BVAR:
- SSVS prior:
set_ssvs() - Horseshoe prior:
set_horseshoe() - NG prior:
set_ng() - DL prior:
set_dl()
- Minneosta prior
-
contem_spec: Contemporaneous prior specification. -
cov_spec: Covariance prior specification. Useset_ldlt()for homoskedastic model. -
include_mean = TRUE: By default, you include the constant term in the model. -
minnesota = c("no", "short", "longrun"): Minnesota-type shrinkage. -
verbose = FALSE: Progress bar -
num_thread: Number of thread for OpenMP- Used in parallel multi-chain loop
- This option is valid only when OpenMP in user’s machine.
Stochastic Search Variable Selection (SSVS) Prior
(fit_ssvs <- vhar_bayes(etf_train, num_chains = 1, num_iter = 20, coef_spec = set_ssvs(), contem_spec = set_ssvs(), cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun"))
#> Call:
#> vhar_bayes(y = etf_train, num_chains = 1, num_iter = 20, coef_spec = set_ssvs(),
#> contem_spec = set_ssvs(), cov_spec = set_ldlt(), include_mean = FALSE,
#> minnesota = "longrun")
#>
#> BVHAR with SSVS prior + SSVS prior
#> Fitted by Gibbs sampling
#> Total number of iteration: 20
#> Number of burn-in: 10
#> ====================================================
#>
#> Parameter Record:
#> # A draws_df: 10 iterations, 1 chains, and 90 variables
#> phi[1] phi[2] phi[3] phi[4] phi[5] phi[6] phi[7] phi[8]
#> 1 -0.12652 0.33810 -0.4064 0.83449 -0.0837 0.0786 -0.06567 0.1567
#> 2 0.14061 0.20717 -0.4043 0.06269 0.1470 0.0134 0.66526 -0.0353
#> 3 0.49999 0.21005 -0.3089 0.03101 0.0705 0.0383 0.20529 -0.0334
#> 4 0.19882 -0.07308 -0.0888 -0.00346 -0.2732 -0.0646 0.00358 -0.0982
#> 5 0.01916 0.00323 -0.1040 0.05931 -0.1829 -0.1766 -0.11164 0.2980
#> 6 0.05360 0.09245 -0.0783 -0.14020 -0.2886 -0.1908 -0.19319 -0.0846
#> 7 0.04501 0.09108 -0.0817 -0.97071 -0.0987 -0.2435 1.25361 1.0924
#> 8 0.24949 0.53449 -0.2133 -2.09922 -0.6911 -0.2955 2.07637 2.0887
#> 9 0.03684 -0.02899 0.0204 -1.24023 -0.2385 -0.4126 1.08892 0.9670
#> 10 -0.00362 -0.07756 0.0123 -1.40019 0.1541 -0.5540 1.78003 1.6721
#> # ... with 82 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}autoplot() for the fit (bvharsp object)
provides coefficients heatmap. There is type argument, and
the default type = "coef" draws the heatmap.
autoplot(fit_ssvs)
#> Warning: `label` cannot be a <ggplot2::element_blank> object.
#> `label` cannot be a <ggplot2::element_blank> object.
#> `label` cannot be a <ggplot2::element_blank> object.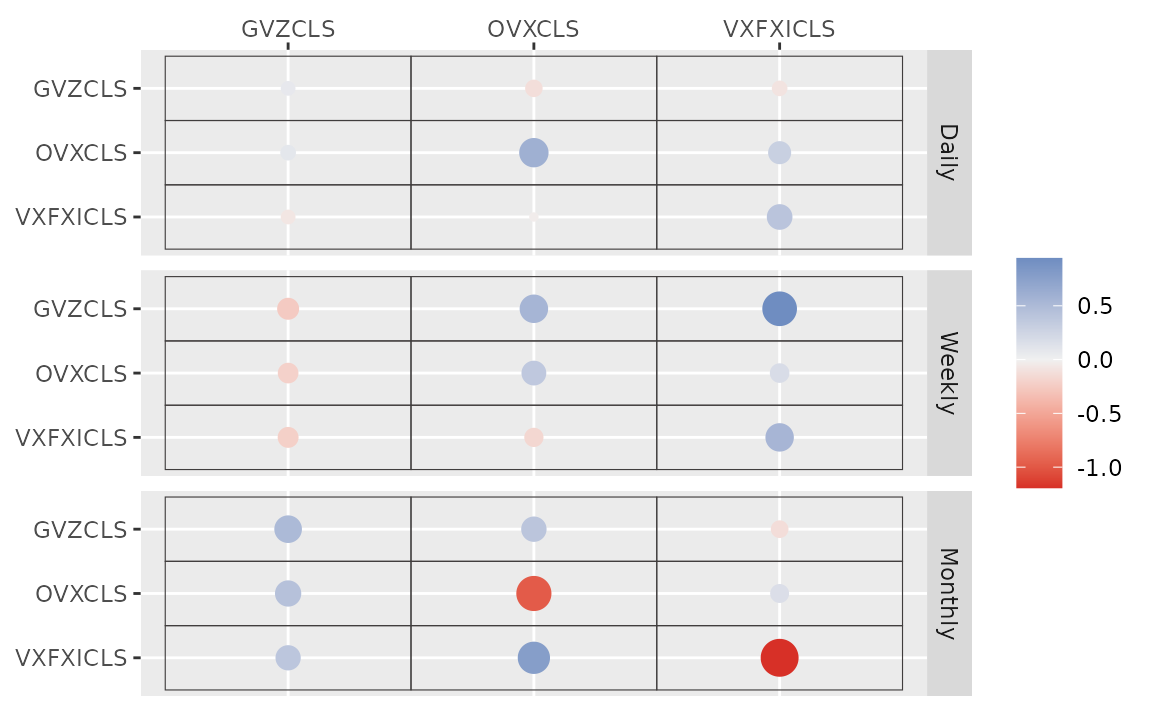
Horseshoe Prior
coef_spec is the initial specification by
set_horseshoe(). Others are the same.
(fit_hs <- vhar_bayes(etf_train, num_chains = 2, num_iter = 20, coef_spec = set_horseshoe(), contem_spec = set_horseshoe(), cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun"))
#> Call:
#> vhar_bayes(y = etf_train, num_chains = 2, num_iter = 20, coef_spec = set_horseshoe(),
#> contem_spec = set_horseshoe(), cov_spec = set_ldlt(), include_mean = FALSE,
#> minnesota = "longrun")
#>
#> BVHAR with Horseshoe prior + Horseshoe prior
#> Fitted by Gibbs sampling
#> Number of chains: 2
#> Total number of iteration: 20
#> Number of burn-in: 10
#> ====================================================
#>
#> Parameter Record:
#> # A draws_df: 10 iterations, 2 chains, and 124 variables
#> phi[1] phi[2] phi[3] phi[4] phi[5] phi[6] phi[7] phi[8]
#> 1 0.3966 -0.04869 -0.073793 -0.11783 0.00293 1.078 0.01429 -0.0789
#> 2 -0.0702 -0.02312 0.102940 0.23771 -0.01322 0.919 0.01128 -0.1104
#> 3 0.0616 -0.16271 -0.022504 0.34424 0.04520 1.022 0.01900 0.0413
#> 4 -0.0212 -0.20881 -0.080155 0.28622 -0.07633 0.978 -0.04185 -0.0055
#> 5 0.0240 -0.21209 0.000571 0.19608 -0.00800 1.055 0.04672 -0.0653
#> 6 0.1203 -0.00323 0.020402 0.00468 0.04875 0.965 0.00531 -0.0131
#> 7 0.0786 -0.06913 0.035241 0.02278 0.02359 1.020 0.00335 0.0411
#> 8 0.0973 0.10813 -0.126033 0.06277 -0.01706 0.887 0.03042 -0.0348
#> 9 0.1122 -0.09499 0.123442 -0.02565 0.00284 0.747 -0.05957 0.0397
#> 10 0.1361 0.00468 0.312354 0.00279 -0.03629 0.854 -0.00382 -0.0222
#> # ... with 10 more draws, and 116 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}
autoplot(fit_hs)
#> Warning: `label` cannot be a <ggplot2::element_blank> object.
#> `label` cannot be a <ggplot2::element_blank> object.
#> `label` cannot be a <ggplot2::element_blank> object.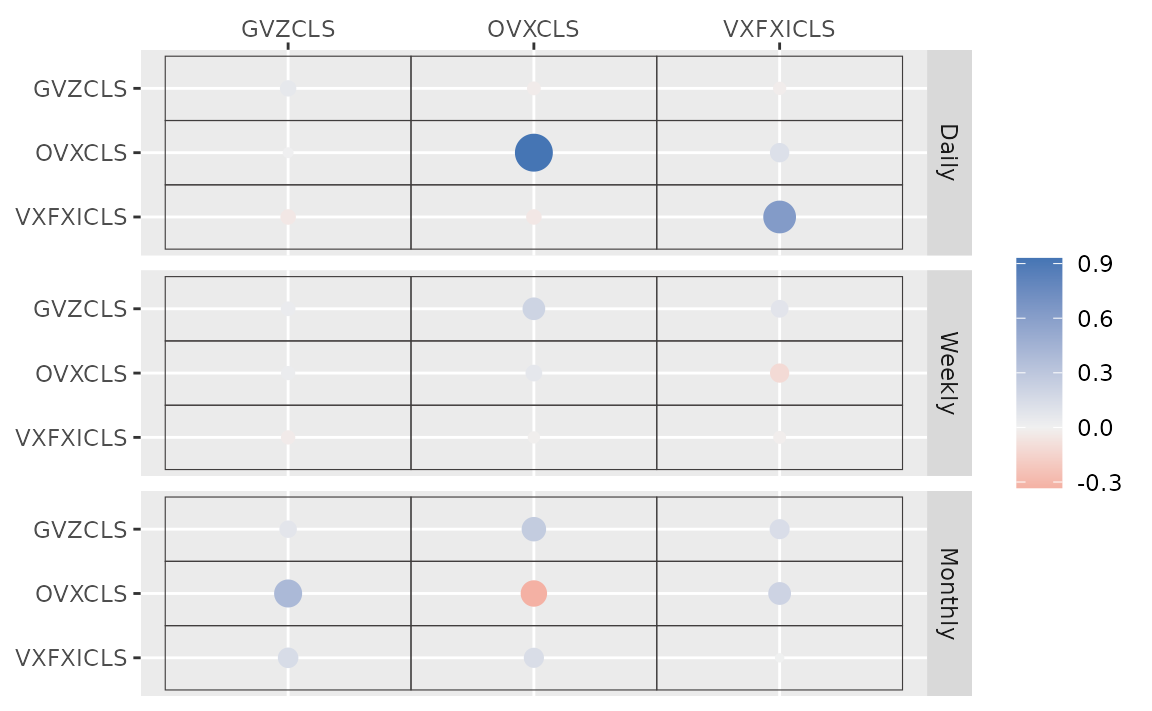
Minnesota Prior
(fit_mn <- vhar_bayes(etf_train, num_chains = 2, num_iter = 20, coef_spec = set_bvhar(lambda = set_lambda()), cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun"))
#> Call:
#> vhar_bayes(y = etf_train, num_chains = 2, num_iter = 20, coef_spec = set_bvhar(lambda = set_lambda()),
#> cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun")
#>
#> BVHAR with MN_Hierarchical prior + MN_Hierarchical prior
#> Fitted by Gibbs sampling
#> Number of chains: 2
#> Total number of iteration: 20
#> Number of burn-in: 10
#> ====================================================
#>
#> Parameter Record:
#> # A draws_df: 10 iterations, 2 chains, and 63 variables
#> phi[1] phi[2] phi[3] phi[4] phi[5] phi[6] phi[7] phi[8]
#> 1 0.4138 -0.0515 0.3069 -0.1077 0.4402 1.101 -0.1993 -0.0312
#> 2 0.3537 -0.2305 -0.0559 -0.1199 0.2445 0.917 0.0434 0.1963
#> 3 0.2705 -0.0465 0.0360 0.0420 0.1887 0.874 0.1031 -0.1930
#> 4 0.3118 -0.1969 -0.0404 0.0694 0.0781 0.823 0.1961 0.0728
#> 5 0.4405 0.0575 0.0506 0.0897 0.0893 0.801 0.1058 0.2704
#> 6 0.0848 -0.2301 0.1407 0.0722 0.1061 1.128 0.2398 0.2817
#> 7 0.1809 -0.0362 0.1832 0.1701 0.0527 1.200 0.3462 0.1840
#> 8 0.1078 -0.2035 0.2829 0.3574 -0.0320 1.036 0.1089 -0.0697
#> 9 0.1737 -0.0848 0.3013 0.2573 -0.0513 0.785 0.0675 0.1181
#> 10 0.2752 -0.1289 0.3415 -0.1696 0.0768 0.940 -0.4074 -0.0673
#> # ... with 10 more draws, and 55 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}Normal-Gamma prior
(fit_ng <- vhar_bayes(etf_train, num_chains = 2, num_iter = 20, coef_spec = set_ng(), cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun"))
#> Call:
#> vhar_bayes(y = etf_train, num_chains = 2, num_iter = 20, coef_spec = set_ng(),
#> cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun")
#>
#> BVHAR with NG prior + NG prior
#> Fitted by Metropolis-within-Gibbs
#> Number of chains: 2
#> Total number of iteration: 20
#> Number of burn-in: 10
#> ====================================================
#>
#> Parameter Record:
#> # A draws_df: 10 iterations, 2 chains, and 97 variables
#> phi[1] phi[2] phi[3] phi[4] phi[5] phi[6] phi[7] phi[8]
#> 1 -0.03575 0.0600 0.0582 0.5228 -0.00426 1.159 0.75115 0.3098
#> 2 -0.01736 -0.0240 0.0191 -0.0935 0.00320 0.989 0.18132 0.1850
#> 3 0.00163 -0.3376 -0.0330 -0.1641 -0.46213 0.795 0.39452 0.0376
#> 4 0.00161 -0.1816 0.1044 0.0895 0.32816 1.007 -0.02902 0.0557
#> 5 -0.00385 -0.1093 -0.0205 -0.3400 0.79470 0.973 -0.01429 -0.0407
#> 6 0.08412 -0.1293 0.0304 0.1529 0.54847 0.894 0.00682 -0.0116
#> 7 -0.05864 -0.0206 0.2211 -0.0567 0.15857 0.506 -0.00174 -0.2649
#> 8 0.13132 -0.2080 -0.0415 -0.0104 0.41555 0.255 -0.02090 -0.3160
#> 9 0.02699 -0.1494 -0.0499 0.1527 -0.24926 0.683 0.06663 -0.2195
#> 10 0.09291 -0.0268 -0.0353 0.2645 0.06679 0.828 -0.02057 -0.0520
#> # ... with 10 more draws, and 89 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}Dirichlet-Laplace prior
(fit_dl <- vhar_bayes(etf_train, num_chains = 2, num_iter = 20, coef_spec = set_dl(), cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun"))
#> Call:
#> vhar_bayes(y = etf_train, num_chains = 2, num_iter = 20, coef_spec = set_dl(),
#> cov_spec = set_ldlt(), include_mean = FALSE, minnesota = "longrun")
#>
#> BVHAR with DL prior + DL prior
#> Fitted by Gibbs sampling
#> Number of chains: 2
#> Total number of iteration: 20
#> Number of burn-in: 10
#> ====================================================
#>
#> Parameter Record:
#> # A draws_df: 10 iterations, 2 chains, and 91 variables
#> phi[1] phi[2] phi[3] phi[4] phi[5] phi[6] phi[7]
#> 1 -0.00240 0.02170 0.000638 -0.494874 0.24782 1.054 -0.06912
#> 2 -0.11518 -0.09074 0.002409 0.288768 0.15906 0.842 0.10505
#> 3 0.09778 -0.05666 -0.009738 0.023870 0.14767 0.794 -0.07326
#> 4 0.10159 -0.04858 0.003269 -0.009243 -0.07123 1.038 0.10176
#> 5 -0.03366 -0.03667 0.003990 0.018902 -0.02215 1.003 0.33110
#> 6 -0.03276 0.00619 0.015556 0.007572 0.40212 0.787 -0.18810
#> 7 0.07294 0.01470 0.026532 -0.001431 0.16105 0.804 -0.00594
#> 8 0.14440 0.05501 -0.079493 -0.000369 0.86305 0.865 -0.00515
#> 9 0.00942 -0.09697 0.112964 0.000574 0.00201 0.985 -0.00477
#> 10 0.05380 -0.05886 -0.033148 -0.000881 -0.00188 0.859 -0.00425
#> phi[8]
#> 1 -4.24e-02
#> 2 -1.26e-02
#> 3 -1.66e-02
#> 4 9.74e-04
#> 5 -4.21e-04
#> 6 -3.26e-05
#> 7 2.61e-05
#> 8 3.38e-06
#> 9 1.05e-06
#> 10 -5.88e-04
#> # ... with 10 more draws, and 83 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}Bayesian visualization
autoplot() also provides Bayesian visualization.
type = "trace" gives MCMC trace plot.
autoplot(fit_hs, type = "trace", regex_pars = "tau")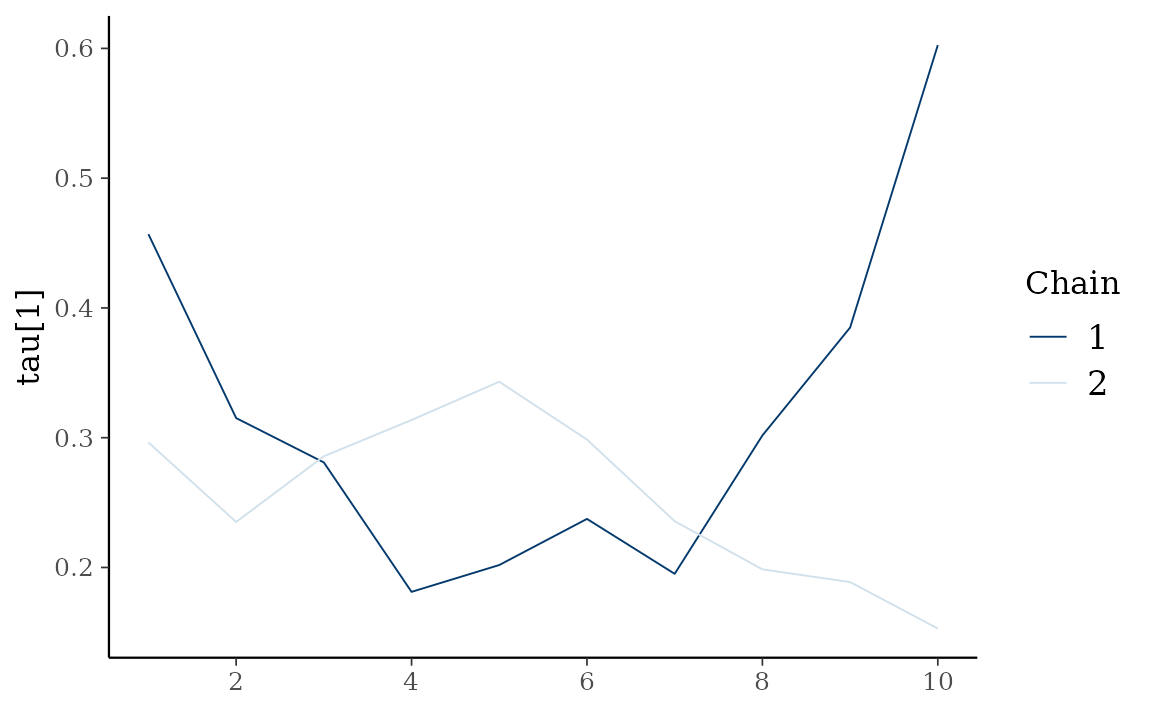
type = "dens" draws MCMC density plot. If specifying
additional argument facet_args = list(dir = "v") of
bayesplot, you can see plot as the same format with
coefficient matrix.
autoplot(fit_hs, type = "dens", regex_pars = "kappa", facet_args = list(dir = "v", nrow = nrow(fit_hs$coefficients)))
